- Cancer diagnosis, biomarker discovery, and multi-omics research
- Research on metabolic diseases and functional biomaterials
- Mass spectrometry-based proteomic analysis and disease mechanism research
- Characterization studies of next-generation biopharmaceuticals
Research Content
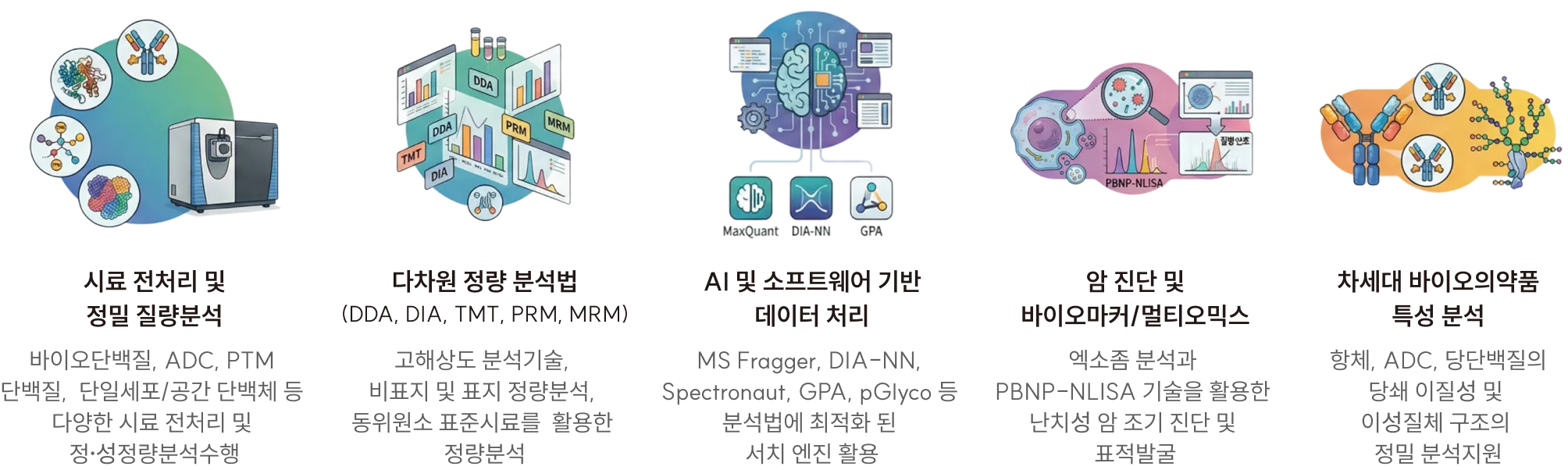
Integrated mass spectrometry platform for next-generation biopharmaceuticals and disease research
- Performs cancer diagnostics, precise characterization of next-generation biopharmaceuticals (e.g., ADCs), and biomaterial marker analysis by integrating high-resolution mass spectrometry with advanced data interpretation technologies
- Maximizes the efficiency and reproducibility of drug development through an integrated platform that connects sample preparation, mass spectrometry analysis, and data processing
- Key data analysis software used across research fields.
| Analysis data type | Software used |
|---|---|
| DDA data | Msfragger, MaxQuant, Proteome Discoverer |
| DIA data | DIA-NN, Spectronaut |
| Glycoprotein | GPA, pGlyco |
| Biopharmaceuticals, antibodies, ADCs | UNIFI |
| PRM & MRM | Skyline |
Cancer diagnosis, biomarker discovery, and multi-omics research
- Applies exosome multi-omics analysis technologies based on proteomics and exosome isolation techniques to develop diagnostic technologies for intractable cancers and to discover and validate cancer targets
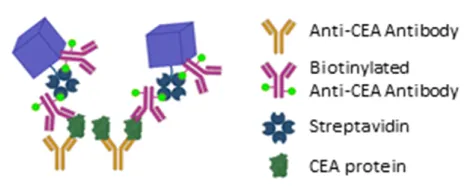
- Development of a high-sensitivity and user-friendly PBNP-NLISA analytical method for early biomarker diagnosis using colorectal cancer and adenoma clinical samples
Journal of Extracellular Vesicles, 2025, Vol. 14 (IF=14.5, top 7% in JCR), June 2025.

Research on metabolic diseases and functional biomaterials
- Performs structural and functional studies by isolating metabolic disease marker compounds from functional biomaterials and analyzing them using MS/MS
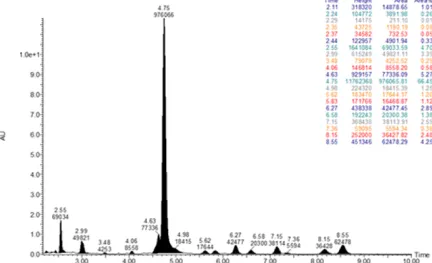
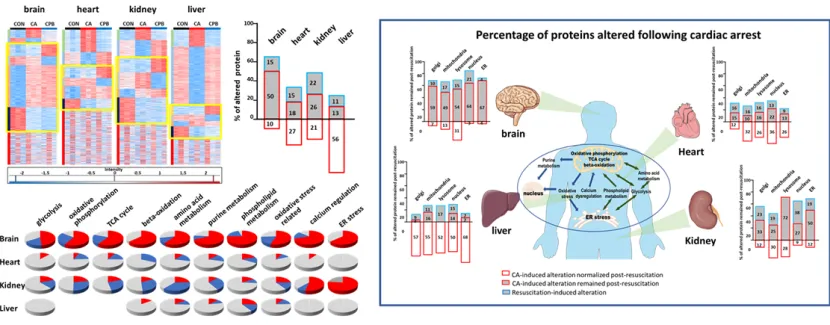
Mass spectrometry-based proteomic analysis and disease mechanism research
- Intracellular proteomic analysis for mass spectrometry-based drug development
- Spatial proteomics analysis using small tissue samples at the single-cell level
- Discovery of biomarkers through qualitative proteomic analysis of clinical samples (blood, tissue, urine) and enhanced quantification using isotope-labeled standard samples
- Investigation of disease mechanisms through mass spectrometry-based analysis of post-translational modifications (phosphorylation and glycosylation) in proteomics
Published in Journal of Pharmaceutical Analysis, Vol. 14 (2024), pp. 427–430 (IF=8.9, top 4.0% in JCR), September 2023
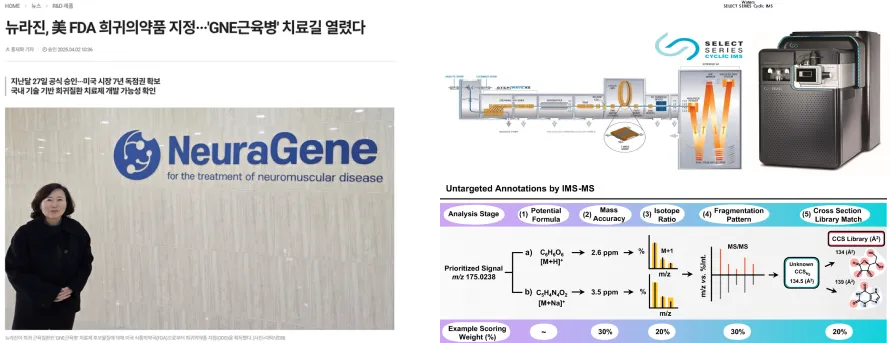
Characterization studies of next-generation biopharmaceuticals
- Characterization of glycan heterogeneity in glycoprotein-based biopharmaceuticals through mass spectrometry analysis
- Advancement of isomer separation and structural characterization of next-generation therapeutics by integrating high-resolution mass spectrometry with multidimensional separation techniques such as ion mobility
Analysis and standardization of NEG-101 required for regulatory approval of a therapeutic for the rare muscle disease GNE myopathy
Research Infrastructure
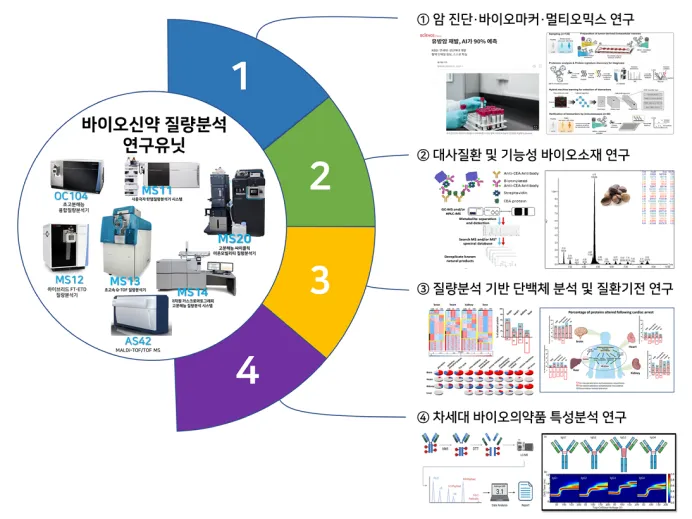
Possesses analytical infrastructure that integrates high-resolution and multidimensional mass spectrometry (LC-MS, MALDI-MS, Orbitrap, Cyclic IMS, etc.) with AI-based data interpretation technologies to enhance precise characterization of next-generation biopharmaceuticals such as antibodies, ADCs, and gene therapies, thereby maximizing the efficiency, precision, and reproducibility of drug development
- The Biopharmaceutical Mass Spectrometry Research Unit (Cho Geon, Choi Jongsun, Jung Youngho, Lee Jooyeon, Yoon Sungho, Kim Minseon) has established an integrated analytical platform supporting biopharmaceutical characterization and proteomics analysis as well as academia–industry–research collaborations, thereby contributing to next-generation therapeutic development and rapid manufacturing while functioning as a global hub for biopharmaceutical analysis
Infrastructure and Research Overview of the Biopharmaceutical Mass Spectrometry Research Unit
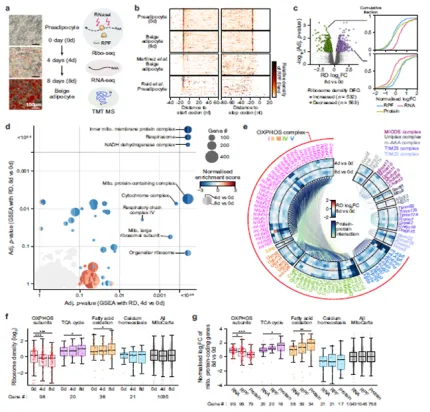
Elucidation that metabolic changes during beige adipocyte differentiation reprogram translational regulation to drive differentiation-specific protein synthesis.
Nature Communications, IF=15.7, top 7.35% in JCR, April 2025
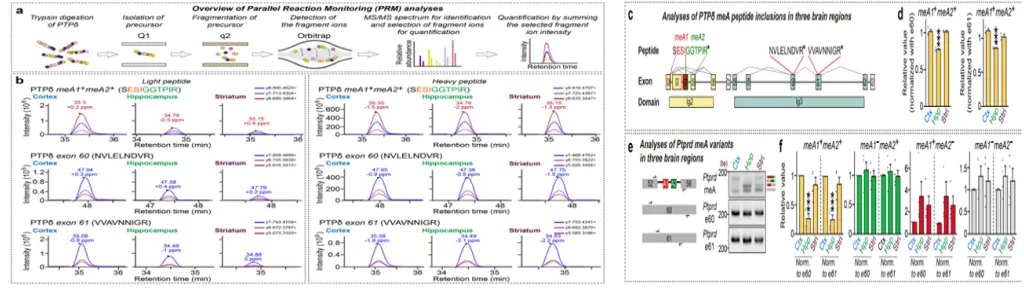
Context-dependent splicing patterns of LAR-RPTP microexons determine circuit-specific synaptic specificity, enabling the elucidation of neural circuit structure and function through proteome profiling.
Nature Communications (2024) 15:1624, IF=15.7, top 7.35% in JCR, February 2024
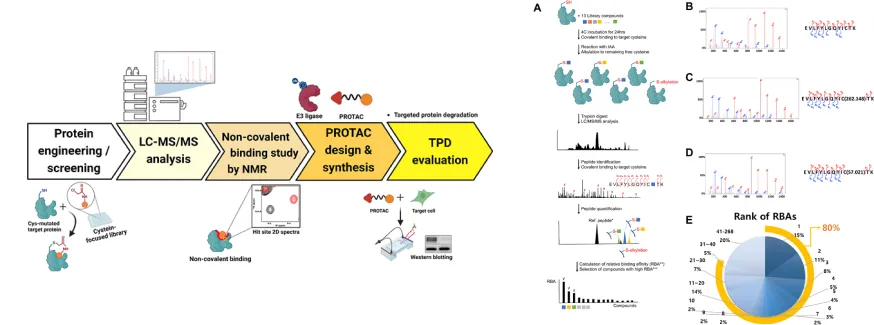
Identification of novel ligands that bind to “undruggable” targets through covalent ligand screening and their application to PROTAC design, suggesting the potential for selective degradation of target proteins (e.g., MDM2).
European Journal of Medicinal Chemistry 263 (2024) 115929, IF=5.9, top 13.9% in JCR, January 2024.
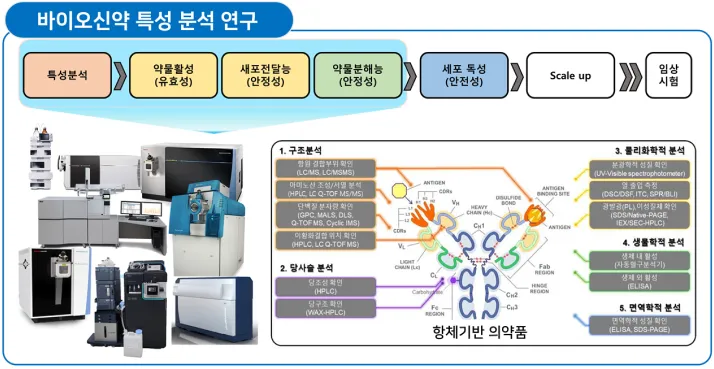
Introduction to characterization research for biopharmaceutical development
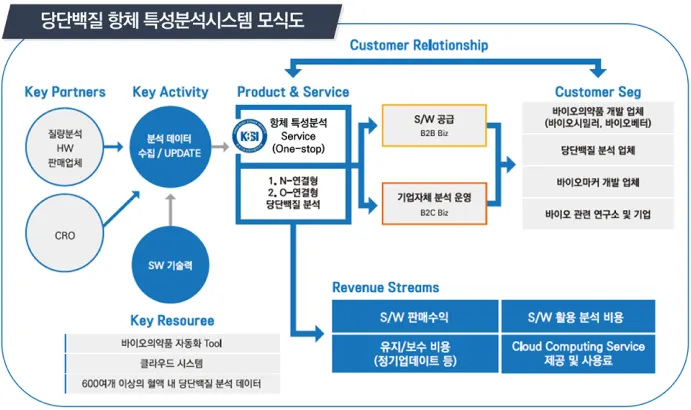
Schematic diagram of the glycoprotein antibody characterization support system
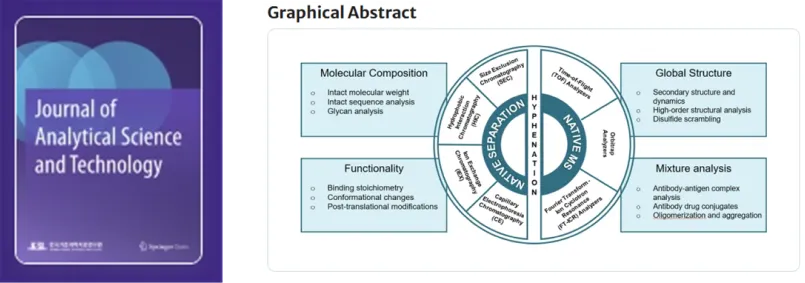
Advances in mass spectrometry-based approaches for monoclonal antibody characterization: structural complexity and analytical challenges.
Journal of Analytical Science and Technology 15, 23 (2024), IF=3.2, top 47.74% in JCR, April 2024. Front cover article